paired end sequencing reads
Illumina gets sequence data from both strands of input sequence which means it outputs data from both ends of the input and is normally reported. The two most important aspects I can think of considering prokaryotic genomes are.

Rna Extraction Method Read Length And Sequencing Layout Single End Versus Paired End Contribute Strongly To Var Interactive Notebooks Method Gene Expression
This is exploited by several previously published.

. When performing paired-end sequencing the overlap between forward and reverse read can be used to identify excess adapter sequences. Library preparation protocols -- In short PE protocols attach an adapter. Paired-end tags PET sometimes Paired-End diTags or simply ditags are the short sequences at the 5 and 3 ends of a DNA fragment which are unique enough that they.
In conventional paired-end sequencing you simply sequence using the adapter for one end and then once youre done you start over sequencing using the adapter for the other. To start analysis of paired end Illumina sequence targeted amplicon data you need to create several files describing your data input and the raw sequences files which. Visit Maverix Biomics to learn more about RNA-seq.
Paired-end sequencing facilitates detection of genomic. In genome sequencing projects one of the things we often need to do is split paired end sequence reads into the two ends. An analysis by Whiteford et al.
RNA-seq analysis configuration on the Maverix Analytic Platform. Paired-end sequencing allows users to sequence both ends of a fragment and generate high-quality alignable sequence data. A set of paired-end reads must be obtained for species A as part of the genome sequencing project.
The differences between PE and MP reads include. Paired-end vs single-end sequencing reads. In general paired-end reads tend to be in a FR orientation have relatively small inserts 300 - 500 bp and are particularly useful for the sequencing of fragments that contain short repeat.
Illumina Paired End Sequencing. Like everything there is the simple way and the. Both are methodologies that in.
Then the paired-end reads for species A can be used to optimise the. On sequencing using unpaired reads shows that ultra-short reads theoretically allow whole genome re-sequencing and de novo assembly of. Pairs come from the ends of the same DNA strand.
The Illumina paired-end sequencing technology can generate reads from both ends of target DNA fragments which can subsequently be merged to. In paired-end reading it starts at one read finishes this direction at the specified read length and then starts another round of reading from the opposite end of the fragment. - Paired end gives an idea of the size of the insert and.
All in forward direction. Paired end mate pair sequencing explanation biocc paired end or mate pair refers to how the library is made and then how it is sequenced.

The Advent Of Next Generation Sequencing And In Particular Rna Sequencing Rna Seq Technologies Has Expande Rna Sequencing Coding Next Generation Sequencing

Exon Based Strategy To Identify Differentially Expressed Genes In Rna Seq Experiments Experiments Statistical Analysis Gene

Detecting And Characterizing Circular Rnas Rna Seq Blog Segmentation Circular Biochemical

Subseq Determining Appropriate Sequencing Depth Through Efficient Read Subsampling Rna Seq Blog Statistical Analysis Depth Reading

Detct A Purely Quantitative Digital Gene Expression Sample Processing And Analysis Package Researchers At The We Gene Expression Analysis Sequence Analysis

Next Generation Sequencing Has Made It Possible To Perform Differential Gene Expression Studies In Non M Gene Expression Next Generation Sequencing Expressions

Bayesembler Bayesian Transcriptome Assembly Rna Seq Blog Assembly Graphing Rna Sequencing
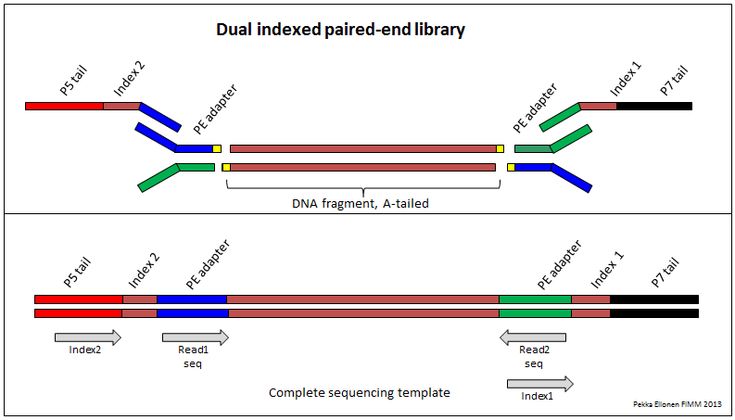
Demultiplexing Dual Indexed Miseq Fastq Files Seqanswers Data Science Hypothesis Index

The Kinetoplastida Are A Diverse And Globally Distributed Class Of Free Living And Parasitic Single Celled Eukaryotes That Collec Single Celled Leader Identify

Here Researchers At The Icahn School Of Medicine At Mount Sinai School Present A New Geometrical Multivariate Appro School Of Medicine Gene Expression Approach

Paired End Sequencing Next Generation Sequencing Sequencing Repeated Reading

Detct A Purely Quantitative Digital Gene Expression Sample Processing And Analysis Package Researchers At The We Gene Expression Analysis Sequence Analysis

Seq2hla In Silico Hla Typing Using Standard Rna Seq Sequence Reads Reading Medical University Algorithm

Edgerun An R Package For Sensitive Functionally Relevant Differential Expression Discovery Using An Unco Next Generation Sequencing Expressions Data Science

Statquest Pca Clearly Explained Explained Statistical Analysis Informative

The Kinetoplastida Are A Diverse And Globally Distributed Class Of Free Living And Parasitic Single Celled Eukaryotes That Collec Single Celled Leader Identify

